Research in the Kurtzman lab focuses on the development of computational methods that aid in the discovery and rational design of new drugs. Our approach applies a combination of statistical mechanical theory and computer simulations to better understand the physical principles governing the molecular recognition between proteins and small molecule drug candidates. His research contributions provide a framework to account for and quantify the role that water plays in molecular recognition.
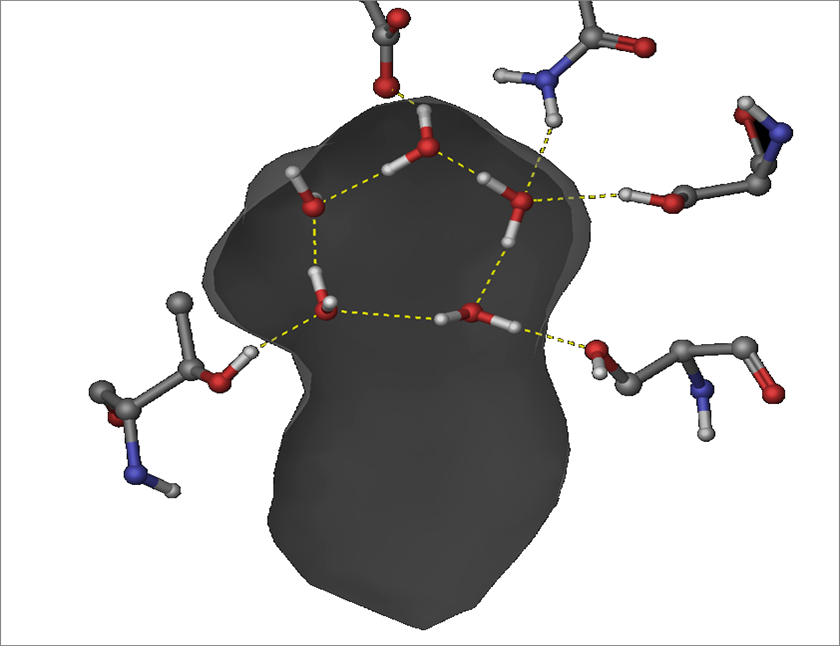
De Novo Drug Discovery
Our approach to de Novo drug discovery is to use information provided by the solvation of both active and allosteric protein sites to direct and aid the search for chemical compounds that can serve as drug leads.
This approach provides several key advantages over structural and ligand based approaches: First, chemical space not explored by these more conventional methods can be directly explored.
Second, no knowledge of binding ligands.
Third, solvation thermodynamics of active and allosteric sites can be used to aid in the search of compound databases and in the ranking of hits.
This last advantage can also be integrated into the more conventional ligand and structure based methods in order to enhance efficiency of these approaches.
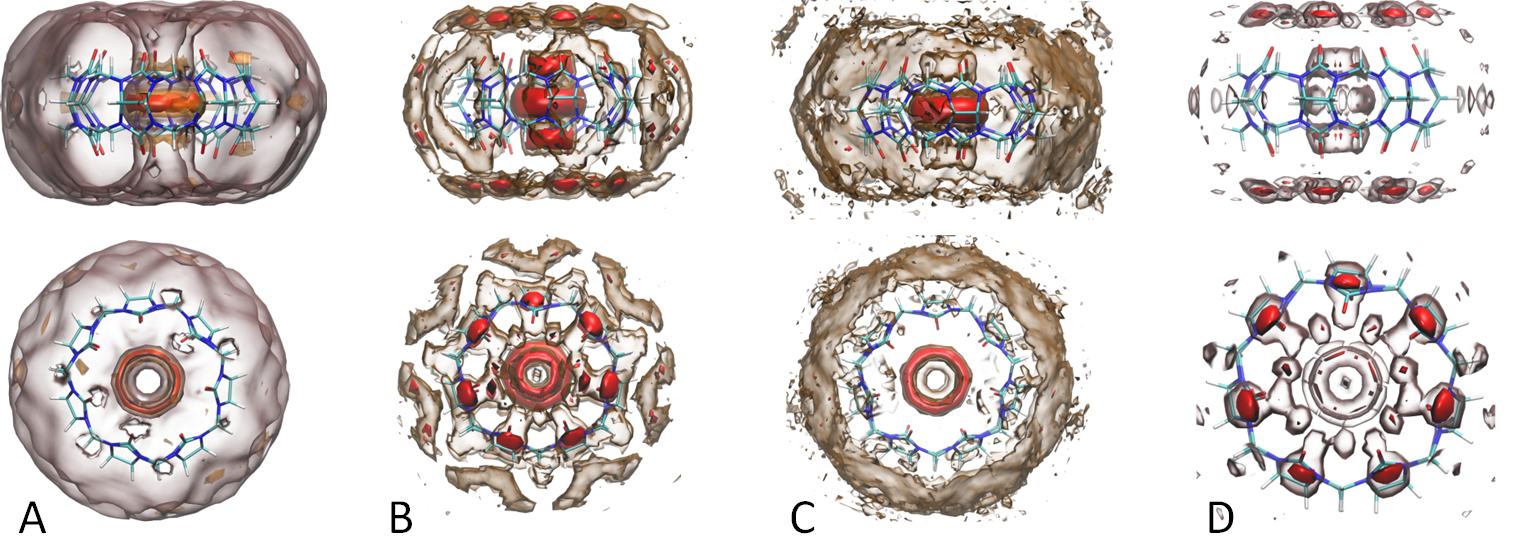
Rational Lead Drug Design
Our approach to rational lead drug design is to utilize solvation thermodynamics information provided by our in-house solvation analysis tools
(GIST & WaterMap) to direct the modification of lead compounds so that they bind with higher affinity and/or specificity for given bio-molecular targets.
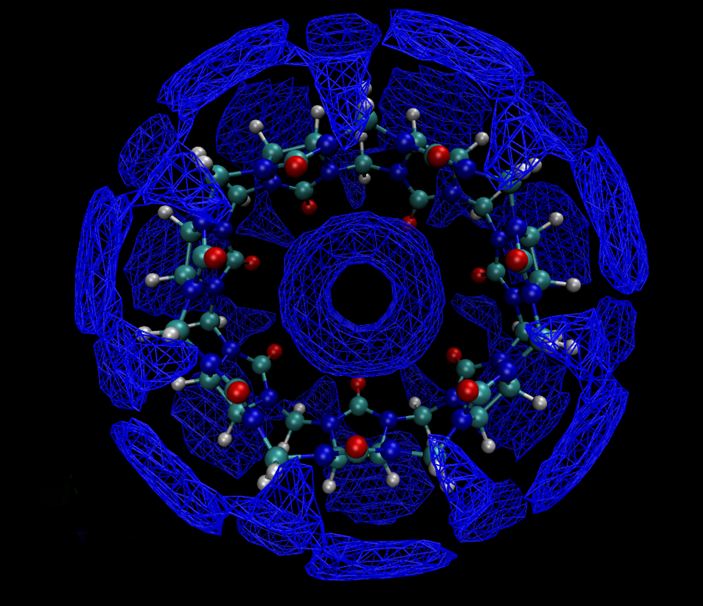
Solvation Analysis Tools
The development and application of computational methodologies that can extract solvation thermodynamics information on a molecular length scale is the main focus of research in this lab.
A significant part of this effort involves exploring how localized solvation thermodynamics can be effectively utilized in a drug design setting.
Two principal methods based on Inhomogeneous Solvation Theory (IST) have been developed by the PI and his collaborators.
The first, a cluster based hydration site approach, has been patented and developed into successful commercial software (WaterMap) that is sold to major drug development companies as well as academic institutions.
A newer, more advanced approach called Grid Inhomogeneous Solvation Theory (GIST), discretizes the equations of IST onto a three dimensional grid and provides a more formally rigorous platform for biomolecular solvation thermodynamic analysis.
The application for localized solvation thermodynamics in the drug design setting alon are manifold.
A number of the applications that exploit solvation thermodynamic information are being applied in our lab including:
Rational lead drug design, De Novo Drug Discovery, Scoring Function Optimization, Allosteric site identification.
